NatureMetrics Expands with Norwegian Cruise Line for 2026 Alaska Season
eDNA water sampling meets guest education — how a science-based monitoring programme is building a longitudinal picture of biodiversity along Alaska's Inside Passage.

The NatureMetrics GCN team has once again scored a perfect ten out of ten in the FAPAS 2019 GCN eDNA proficiency test, one of only two companies that were able to achieve this. In addition, no sample degradation was detected in any of the NatureMetrics kits.
This blind test consisted of 10 samples (A – J) prepared using each company’s own kits. The samples comprised:
Eight companies took part in this year’s proficiency test, and the results are summarised below. NatureMetrics was laboratory 3.
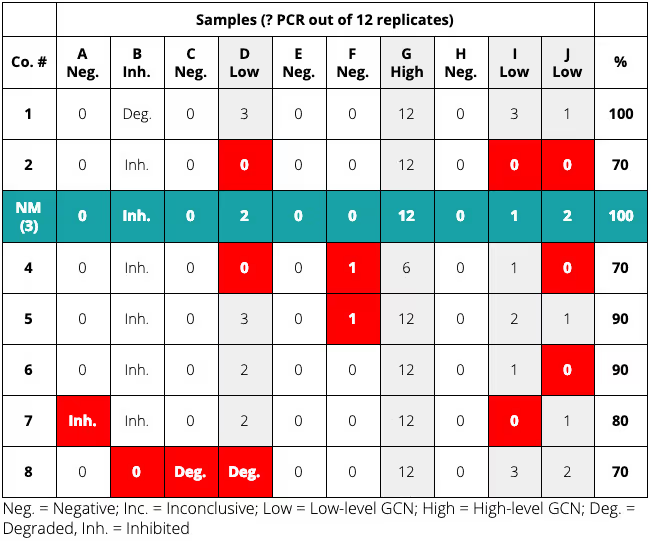
NatureMetrics and Company 1 correctly detected GCN in the one high-level and in the three low-level GCN samples, correctly returned an inconclusive result for the one inhibited sample, and correctly returned negative results for the six blank samples.
All of the companies returned positive detections for the one high-level GCN sample, but only NatureMetrics and Companies 1 and 5 returned positive detections for all four GCN samples.


Companies 2, 4, 6, and 7 returned false-negative results by failing to correctly detect GCN in one or more low-level GCN samples, and Company 8 returned a false-negative result for sample B, which should have been reported as inconclusive (due to inhibition, see below). False-negative results give the client a false sense of security and might mean that positive GCN ponds are not properly safeguarded.
When the level of GCN is low, the effect of stochasticity is higher because the number of positive replicates is low. For example, the average number of positive replicates among the three low-level GCN samples in this test was 1.25 out of 12 possible (range 0 – 3) versus an average of 11.25 positive replicates for the one high-level GCN sample (7 out of 8 companies reported 12 positive replicates and one reported only 6 positive replicates). It is possible that additional replicates would have returned a positive result. A deeper analysis of the interpretation of low-level positives is here.
Companies 4 and 5 returned false-positive results for sample F. This type of error is avoidable with good laboratory practice. The contamination could have been introduced at any stage in the process from sample lysis, DNA extraction, or qPCR setup. One possibility is that sample G (high-level GCN) contaminated the adjacent sample F (a negative) while both tubes or wells were open at the same time.
Because all positive detections require additional population-level assessment by the client, a false-positive eDNA result incurs unnecessary and costly revisits to the water body. Revisiting true-positive GCN sites is already logistically challenging, given that half the site visits have to be completed by mid-May.
Companies 7 and 8 returned false-inconclusive results when the results should have been reported as negatives. False-inconclusive results have implications for the client because returning an inconclusive result when the sample should have been reported as a negative will incur additional costs as the client has to do additional sampling and testing. There are two ways in which a sample can cause an inconclusive result: (1) inhibition and (2) degradation.
(1) Inhibition means that the water body contains high levels of substances, such as humic acids from rotting leaves, that inhibit detection of GCN DNA. Inhibition was reported by Company 7 for sample A, which was, in fact, a true negative.
(2) Degradation means that the kit might not have successfully preserved GCN DNA, which is checked by confirming that the control DNA, which is added to the sample tubes before shipping to the client, can be detected after the kit is returned. Companies 1 and 8 reported that the control DNA in one or more of their samples was degraded (i.e. could not be detected).
None of the NatureMetrics test samples showed evidence of degradation, and NatureMetrics also correctly reported inhibition for test sample B, which was the sample that had been designed to induce inhibition.
For existing customers you can now log in to your account at my.naturemetrics.co.uk and start placing your orders online. From the 2nd March our new ordering system will be going live, this system is more intuitive and easier to use with a fresh new look. This won’t affect your current accounts and any orders placed during January-February will be transferred over. Orders placed before 12 pm will be delivered to you the next working day – although advance notice is appreciated whenever possible. For accounts, ordering, & price enquiries, please contact our team at gcn@naturemetrics.co.uk. Also, please get in contact if you have large orders exceeding 100 kits, for a bespoke quote.
Our NM sampling app allows you to log and submit field data on site. The app is a great way to ensure your samples are processed as efficiently as possible and can even reduce the turnaround time.
We will be very happy to visit you and talk through how to use the kits and explain the process in the lab and how we analyse the results. We can also demonstrate how to use the online system.
.jpg)
eDNA water sampling meets guest education — how a science-based monitoring programme is building a longitudinal picture of biodiversity along Alaska's Inside Passage.
